

There’s something about Listeria that makes it a food safety nightmare. The bacterium has caused thousands of infections over the last few years with an alarming case fatality rate of 21%, primarily in the elderly. It can survive a in a number of environments, including low temperatures, acidic conditions and even high salt. Then there’s the rather problematic production of biofilms, which can resist disinfection. All told, the pathogen has become one of the most important – and as a result studied – foodborne threats.
Over the last 30 years, research has revealed Listeria has a collection of virulence factors necessary to cause infection. One of these is known as Listeria Adhesion Protein (Lap), which allowsadherence to human intestinal cells and signals the body to move the bacterium to the organs. In 2010, a variant of Lap, denoted as LapB, was identified as a factor in maintaining attachment to cells as well as aiding intracellular invasion. In essence, LapB was a critical component needed to infect.
Further research into LapB at the genetic level revealed its code for protein formation, mRNA, is highly protected when the bacterium is encountering environmental stress, particularly during a fight with the immune system. The bodyguards are comprised of proteins, known as Hfq (from the E. coli Host factor for phage Qβ RNA replication) and small RNAs (sRNA) denoted Lhr (for Listeria Hfq-binding RNA).
There are three types of Lhr, denoted LhrA, LhrB and LhRC. Each one has a specific mechanism in the process of bringing Listeria’s genes to protein life. Both LhrA and LhrB have only one copy made but LhrC has 5 different variants. At the time of their discovery in 2006, the function had yet to be elucidated although there was some belief the altered structures had some environmental significance.
The hunt went on for eight years but now the answer appears to have been found. Last week, the same Danish group who discovered the Lhr sRNAs revealed why LhrC has so many different faces. The answer lies in the need to ensure LapB is produced during an infection, regardless of the environment. As the authors learned, LhrC provided the ability to deal with any challenge and keep virulence alive.
The team first wanted to find out whether environmental stimuli lead to the production of LhrC. They tried almost everything in the human arsenal – beta-lactam antibiotics, bile salts, high salt solutions, acid and even alcohol. In all cases, LhrC was present. Looking further, the group found two specific environmental response proteins necessary for transcription. The first was a two-component protein known as LisRK (response Regulator + histidine Kinase), which is involved in reacting to environmental stress. The second was CesRK for CEphalosporin Sensitivity response Regulator) which helps the bacterium resist the presence of beta-lactam antibiotics and alcohol.
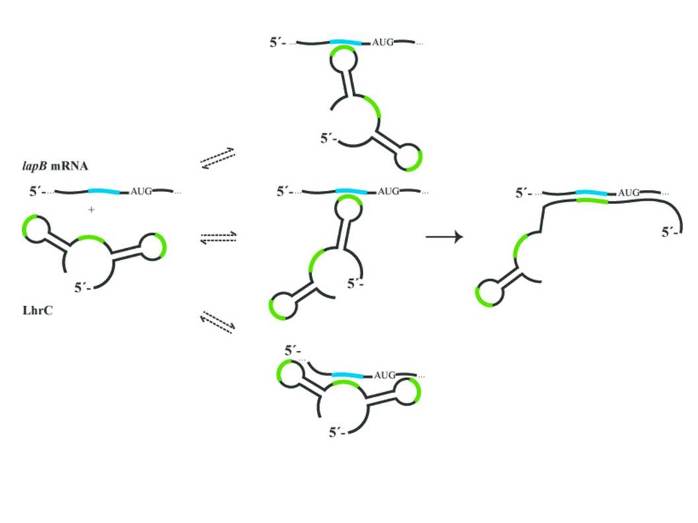
The next step was to determine the target of LhrC. All five variants had an affinity for the LapB mRNA and helped stabilize the mRNA strand for translation into protein. There appeared to be no difference between the five faces in their ability to bring LapB to life. However, while this revealed the function, it did not unveil the reason as to why five copies were needed.
Upon looking even closer at the secondary structures of the LhrC faces, there were few if any differences in the way they appeared. All could have multiple shapes and interact in a number of different ways with LapB mRNA. As to why that might be, the authors needed to go down to the sequence level. There, they found motifs containing the nucleosides cytosine and uridine (CU). Each of the five LhrCs had three different placements for this sequence. Taken together, there were some seventy-five different conformations that could interact with the mRNA and aid translation.
Why this level of redundancy would be needed was not evidently clear. The authors, however, suggested these motifs served as modulators of a butterfly effect. A small ultrasensitive reaction between the CU region and the LapB mRNA could lead to a rapid, large scale reaction to a stressor. No matter what was thrown at the bacterium, it would fight rapidly and effectively.
The results of this work offer perspective as to why_ Listeria_ is so hard to kill in any environment. Unless the bacterium is completely overwhelmed, whether through disinfection, acid treatment or temperature, the chances for survival are good. Moreover, in the midst of an infection, the bacterium can use this system to resist the actions of the immune system and any other treatment such as antibiotics. In effect, the redundancy appears to render the bacterium unstoppable.
For food safety experts, the news is even worse. In order to prevent infections, the highest level of compliance to regulations is needed to ensure the bacterium doesn’t enter the food continuum. Sadly, lapses happen regularly and outbreaks are becoming more common. This means an even greater responsibility is put on the public to be extra careful with any food supply through proper hygiene, food handling, cooking and cleaning. Without adhering to the rules of food safety, you may have a rather nasty encounter with Listeria and a one-in-five chance of not making it at all.
