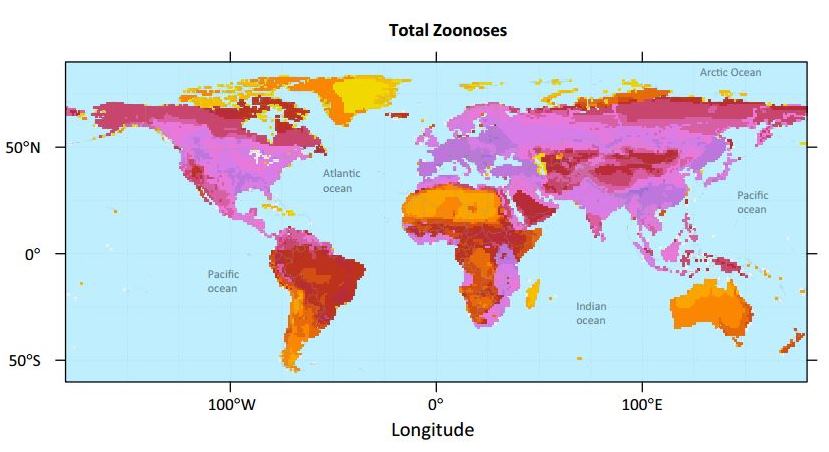

The last few years have seen a rise in rare and unknown infectious diseases. The Ebola epidemic may be the most prevalent example in recent times other pathogens such as MERS, avian and pandemic influenzas, and Nipah virus have emerged in human populations. While the causes of these diseases are quite different in nature, they do have one thing in common. They all originated from animals.
Infectious diseases from animals are known as zoonotics and they are far more common than most realize. Up to 70% of all human infections originates from an animal source. The highest risk comes from animals used in agriculture yet, wild animals also pose a threat through interaction with human populations.

The rise in the number of these zoonotic infections has led some researchers to question whether current public health measures are still useful for certain diseases. In the case of Ebola, the measures were mainly reactionary as the outbreak had gripped West Africa long before appropriate actions could be taken. For some, the absence of an early-warning signal through surveillance of mammals allowed the epidemic to become far worse than it should have.
The concept of looking out for infections in animals is a useful one however logistically, it can be a nightmare. Tracking the various species on a global scale is simply not possible in light of the human and economic resources available. In order to truly have a process that works, a more focused approach is needed.
Last week, an American trio of researchers released a discussion paper on the issue of tracking mammals on the global scale. The hope was to explore the potential for zoonotic infections in different areas of the world and lay out a foundation for prediction and prevention of infectious diseases, outbreaks, and epidemics.
Based on the article, the best approach is to identify disease hotspots where the risk is greatest for transmission between mammals and humans. For example, mixtures of human and bat populations could be considered a hotspot if Nipah virus was found in the animals. The same could be said for areas where humans and porcine species co-exist as they can readily carry a number of human pathogens. These environments would see frequent opportunities for transmission and increase the risk for an outbreak.
While these two examples are well-known thanks to historical information, determining other hotspots may not be as easy. The prime example in this case was West Africa in which Ebola was not expected to cause any significant troubles due to a lack of any significant interaction between humans and wild mammals. To determine hotspots, an evaluation of all possible zoonotic interactions is needed.
This is, however, not an easy task. As the authors reveal, there are 27 terrestrial mammal orders and large species richness. Tracking all the species even in a designated hotspot area would be difficult at best. The only way to approach this would be to examine those with at least some potential interaction with humans. Specific orders of highest risk would include rodents, bats, and carnivores. This may seem obvious but that is only due to the emergence of potentially preventable outbreaks.
The final perspective in developing a foundation is the nature of the pathogen. Infectious diseases affect mammals differently and outcomes vary significantly. Although in the case of lethal pathogens such as Ebola, MERS, and plague, there is little doubt as to their importance, other pathogens may pose equal risk even if they are not as deadly.
One particular example not mentioned in the paper is hepatitis E virus (HEV). The infection is not usually fatal yet can be a serious threat to pregnant women as in India, up to two-thirds end up succumbing to the disease. The route of transmission isn’t direct contact with live animals but through the ingestion of improperly cooked meat. Based on the potential severity of the disease, any area with a high prevalence of the virus in an animal population would be a hotspot.
Based on the information presented in the article, identification of hotspots might be the best route moving forward. While it may not allow us to catch every possible zoonotic transmission – they still occur in coldspots – the practice may allow us to predict possible outbreaks. In addition, through mandated changes in human behaviour, outbreaks may even be prevented. In consideration of the impact of some recent zoonotic epidemics, this approach may be well worth continuing and enacting.
