

Meet Saccharomyces cerevisiae, the humble brewer’s yeast and resident co-deity to BeerSci’s brewing endeavors.
Humans have been exploiting S. cerevisiae‘s fermentation prowess for thousands of years. Without it we wouldn’t have beer, bread or wine. In addition to its uses in food production, S. cerevisiae is also an amazing tool for molecular and cell biology, one that is helping scientists suss out the rules of how our cells work and gain clues to what happens at the molecular level when things go wrong.
Much of what we understand about eukaryotic cellular processes and eukaryotic gene expression, we learned by studying S. cerevisiae.
That’s because S. cerevisiae is one of the simplest eurkaryotic cells–cells like those that make up your dog, your houseplants or your local bartender. In fact, in 1996 S. cerevisiae became the first eukaryote to have its genome sequenced. According to the Saccharomyces Genome Database, S. cerevisiae‘s genome has some 12,100,000 base pairs and some 6,600 open reading frames (that is, places in the genome that could possibly contain a gene).
Most of you, I am sure, remember that there are two general kinds of cells: prokaryotic and eukaryotic. That is, “no nucleus” and “has a nucleus.” That’s all true, but the differences between the two kinds of cells are much more profound than that. Bacteria — prokaryotes — organize their genetic material in a completely different (and much simpler) way than do eukaryotes. Prokaryotes usually only have a chunk of DNA for a genome — usually circular — and a few extra chunks, called plasmids, kicking around in the cytosol. Those plasmids are really useful in doing things like sharing genes between bacteria, and its how one antibiotic-resistant strain of bacteria can pass along antibiotic resistance to a bunch of nigh-unrelated strains of bacteria in, say, your intestines. The genes in bacteria are generally read exactly as they are found in the DNA, kind of like how you’re reading this sentence. No intervening clumps of letters to clutter things up.
Eukaryotes, on the other hand, bundle up all that DNA (and they have a lot of it) into a protein-DNA complex called chromatin, then wind that chromatin into individual chromosomes. Further, the genes are constructed in such a way that they must be heavily processed before they can ever “code” for a functional protein. Much of what we understand about eukaryotic cellular processes and eukaryotic gene expression, we learned by studying the molecular mechanics of S. cerevisiae.

Why am I going on about all of this? /Because when I was researching the S. cerevisiae genome to find out which gene loci or other factors are responsible for, say, the ester-producing qualities of British ale yeast or the heat-tolerance and high-attenuation profiles of saison yeasts, I found something so cool I wanted to write about it. I found an article from the 20 May 2010 issue of Nature that described how biologists and computer scientists at the University of Washington had determined how chromosomes might be arranged in a normal yeast cell and how that organization might affect gene expression. Eventually, this could have implications in how we understand human gene expression.
A bit of background: One of the most surprising findings of the last decade or so is that it’s not just what’s on the genes that affects an organism’s physical and chemical makeup, but how those genes are packaged in chromatin that matters. For a field that thought for over a century that genes are king, that’s a crazy result, and one of my favorites in recent memory.
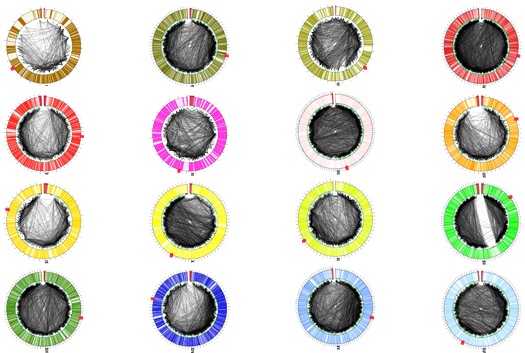
But it also turns out that not only does the gene packaging matter, so does the chromosome placement in the nucleus. Think of it this way: if you had a Xerox machine and a certain amount of time, chances are that you’d Xerox any documents that were closest to you a lot, rather than drag that machine across the office to copy something far away. That’s what chromosome organization does–it doesn’t matter if two genes are somewhat near each other on a linear strand of “genome” if they end up halfway across the nucleus from each other. Instead, it’s also important if areas on two totally different chromosomes get cozy next to each other in the nucleus–you get a lot of copies of gene product from chromosome regions that are adjacent to one another. The paper in question detailed how the researchers were able to look at all of the spatially adjacent chromosome loci by using a relatively new method to create DNA libraries of those interactions, then map and model how the 16 chromosomes in the S. cerevisiae nucleus interacted with each other.
I spoke with one of the paper’s authors, William Noble, about the method and the results. One of the more surprising (and initially worrying) results of the work was how chromosome XII differs from the other chromosomes. The data showed that, unlike the others, XII had a huge swath that interacted with hardly anything else. Noble said that he and his colleague who ran the analysis worried that there was an artifact in the data, making chromosome XII look different when it wasn’t. But further scrutiny uncovered the answer: Unlike all of the other chromosomes, XII sits partially in a large structure called the nucleolus, where ribosomal RNA is made. The nucleolus acts like a barrier so the arms of the chromosome can’t get close to each other–kind of like how it’s hard to give a sumo wrestler a bear hug without Inspector Gadget extendable arms.

The researchers also discovered that most interactions between chromosomes happened at sites adjacent to the centromeres. And, perhaps not surprisingly, the sites known to code for tRNA, no matter where they were found on a chromosome, tended to associate into clusters, and that one such cluster localized to the nucleolus. It makes sense that tRNA genes would cluster in an area where active translation and protein synthesis happens.
After we chatted about the findings of the paper, Noble and I talked about the possible implications. Other studies have suggested (and this one reinforces them) that nuclear ordering–how chromosomes organize themselves in the nucleus–actually does affect how genes are expressed in an organism’s day-to-day existence. The story behind “are you a product of your genes” is becoming murkier by the month. Similar studies have been done for other genomes, including human, but because those genomes are large and complex, it’s not possible to get the same resolution the University of Washington group got with budding yeast.

None of this has anything to do with brewing, I know. But just like falling down the wiki hole sometimes leads to interesting discoveries, digging through dozens of scientific papers on the yeast genome leads to some other interesting discoveries.
To bring this back to brewing again, here’s a recipe similar to one for an ale brewed around the same time that European scientists rediscovered Gregor Mendel’s work on “discrete units of inheritance.” We modeled it after the November 15, 1901 KK by Pretty Things Ale Project, itself based on a recipe for a beer brewed in London on its namesake day. It’s a partial mash recipe because Team BeerSci doesn’t have the equipment to hit a target gravity of 1.079 in a 5-gallon all-grain batch.
INGREDIENTS
4 pounds Maris Otter
1 pound brown malt
10 oz Crystal 120
6 oz Crystal 80
1 oz chocolate malt
4 pounds extra light DME
2 pounds invert no. 3
Add 2.5 pounds of the DME at flameout to keep boil gravity down and increase hop utilization.
1.5 oz East Kent Golding and 0.15 oz Centennial — 90 minutes
1.5 oz East Kent Golding and 0.5 oz Bramling Cross — 60 minutes
1.5 oz East Kent Golding and 1.5 oz Bramling Cross — 30 minutes
Wyeast London III (1318)
INSTRUCTIONS
Rack after 2 weeks and dry-hop in the secondary with 1 oz. of EKG and Bramling Cross apiece.
The beer ended up lighter than the November 15, 1901 KK ale that served as inspiration for this one (more of a chocolate brown than dark brown), but it was still a nice winter ale at 8.1% ABV.
Next week: Barley genome decoded!
Follow BeerSci on Twitter! @BeerSci
