
A new “find-and-replace” genome editing method enables scientists to make large-scale changes to the genetic code of a living cell, faster than previous editing technology by a factor of two. The new method could be used to engineer cells that produce new proteins, or to design genetic “firewalls” that would prevent engineered cells from spreading their DNA.
Using a new genome engineering method, researchers from Harvard University and MIT were able to systematically replace one three-letter DNA “word,” or codon, with another one, throughout the entire genome. Their method reimagines genomes as editable templates, allowing entire sections to be rewritten without interrupting the overall organizational structure.
Ever since genome sequences were unraveled, scientists have been experimenting with ways to edit them, to give cells new capabilities and to cure diseases. But progress has been slow, in part because the editing job must be incredibly precise — one misplaced allele on a chromosome can be lethal. Introducing individual changes is therefore laborious and difficult.
Genetic scissors like zinc-finger nucleases can do the job, glomming on to certain sequences of a genome and cutting it out. Engineered viruses can then bring new DNA to the targeted area. Just last month, researchers successfully snipped out the genes that cause hemophilia, in a study involving living mice.
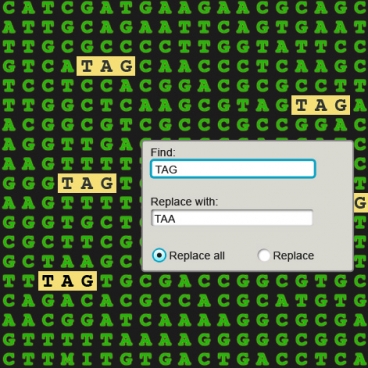
Rather than using scissors, this new method is akin to the find-and-replace function on your word processor, seeking out individual codons and replacing all of them with something new. The researchers used E. coli bacteria and edited a codon that only appears 314 times in the genome, spelled TAG.
Most codons spell out the sequence of an amino acid, as a Harvard Medical School news release explains. But some codons direct a cell to stop adding amino acids to a protein chain. TAG is one such stop codon, and it happens to be the rarest word in the E. coli genome. The team used a method called multiplex automated genome engineering, or MAGE, to replace the TAG codon with another stop codon, TAA.
MAGE locates specific DNA sequences and replaces them with a new sequence as the cell copies its DNA. The team made 32 E. coli strains, each of which had 10 TAG codons replaced. Then they used a new technique called conjugative assembly genome engineering (CAGE), which allowed them to control the process that bacteria use to swap genetic material.

An evolutionary tournament of sorts reduced the number of strains to 16, each with double the amount of TAG edits that it started with; ultimately, a final four strains had one-fourth of all their TAGs replaced, MIT News explains.
The researchers believe they’re on track to replace all 314 of them — without harming the cells in any way.
The next step would be to take out the genetic sequences that allow the cell to read the TAG stop instructions. This machinery could then be used for an entirely new purpose, like encoding a new protein or function.
The research is described in today’s issue of the journal Science.
